FHIR is the HL7’s modern approach to connecting components in the health computing space. Unlike the HL7v2 message approach, FHIR is oriented to enabling applications connect to back-ends. It has been running for a few years now, and is doing good work on how to use REST, terminology, and generally make the application programmers experience better.
It is also over-hyped. That’s not mainly the fault, as far as I can tell, of the core developers but more of an industry exhausted by attempts to make HL7v3 work, but still desperate for solutions. Some of the hype would have you believe that FHIR solves all health informatics problems; common sense says this is not true. A lesser version of the hype would have you believe that FHIR solves all interoperability problems in health. A superficial inspection shows this is also not true. Both versions of the hype are leading some vendors, providers and even some government programmes astray.
My point here is not to criticise FHIR, it’s to encourage those trying to solve the problems of e-health to take a step back, have a good look at the list of problems we know about, and then understand which of those FHIR is going to help with.
This post takes a look at what FHIR’s scope is today, and makes a proposal to significantly extend its capability, in a way that could even help to end the ‘health informatics wars’.
FHIR Scope
In terms of the general scope of health informatics interoperability problems, there are many big questions, including:
- EHR/CDR system design: this requires solving problems of representation of data as they are captured and persisted, e.g. models of what can be recorded; how to code it, also how to query it, export it etc;
- B2B or system-to-system communication: this requires an approach that can shift large amounts of data corresponding to numerous patients, tests or other data entities, normally in the form they were originally captured (i.e. whole data sets);
- C2B or app-system communication: this requires API models of the system such that applications can obtain data and services relevant to their function, and which typically want query views of data, i.e. projections and computations rather than the original data sets;
- Big data and standardised, semantic querying: this requires solving the problem of a standardised semantic approach to marking data, so that in data obtained from different sources, a) basic identification of what the data are about (e.g. BP) is possible and b) things like current BPs are not mixed up with target BPs and diagnosed allergies are not mixed up with exclusion statements for the same allergies;
- Notifications networks: creating and effecting notifications about events to relevant actors requires models of workflows and actor networks.
In the above, FHIR primarily applies to the C2B or app-system communication category. But as we well know, the other categories are equally critical to implementing a running solution, and in some cases, significantly more complex.
FHIR has undoubtedly generated a lot of hype because the ‘app space’ is the currently exciting one, in the era of mobile devices.
However, models of APIs needed for app/system communication don’t exist in a void, but are dependent on the models of content of the EHR as well as models of querying. For example, if your API provides a function
getMedicationList (t: DateTime): MedicationList
(or an equivalent REST GET call + URI) then clearly you need to have defined what a MedicationList is, and this will come from content models.
It is of course technically possible for an API layer to invent its own models of content, effectively imposing them on other systems, but this carries with it implications:
- lack of scalability: we know from existing semantic modelling activities such as SNOMED CT, various guidelines work, openEHR and CIMI that there are hundreds of person-years of work implicated in modelling clinical medicine semantics, as discussed here – an API specification that creates its own definitions is up for replicating such work, which is unlikely to be economically viable;
- semantic mismatch: the API models of content will typically clash with system models of content, necessitating transformations that are inevitably error-prone and potentially performance-degrading.
Can it make sense to try and define all the clinical models yet again as FHIR resources? Well, if the systems you are talking to have no published models of content, as many don’t, then it is potentially the only approach in the short term at least for some key resources. Such systems tend to fall into two categories:
- the organically evolved bowl of spaghetti, typically in-house developed or from small historical suppliers who continue to use a very old core – no architecture or model documentation is typically available for such systems;
- large vendor solutions whose content and query models are kept proprietary as a matter of commercial strategy.
If we were to do a survey of EHR/EMR solutions currently in operation around the world, the majority of instances will fall into one of the above two categories, so there is certainly an argument for an API standard such as FHIR to invent its own content models.
On the other hand, there are many systems that do have explicit architectures and models of content, and this will most likely be the way of the future, as modelling of semantics is essential to intelligent computing functions in the analytics and big data areas. For these systems, connecting to FHIR is potentially difficult, because it requires mapping already well-defined content to independently defined models of content (clinical resources etc) in the API layer which do not typically match.
Where should Domain Semantics be Modelled?
There is another way to see the problem of modelling of domain semantics: as an activity independent not just of products and vendors but also independent of implementation technology standards. This has been the approach of the major terminologies, guidelines, of Intermountain Health with its Clinical Element Models, of various agencies via CDA templates (e.g. VHA), and of openEHR and ISO 13606-based implementers. Semantic models for health arguably fall into 3 major categories:
- terminology and terminology sub-sets;
- models of content – openEHR archetypes, CIMI archetypes, CDA templates etc;
- guidelines.
There are of course many more, e.g. models of ‘study data-sets’ as defined in CDISC’s BRIDG models and also underlying ontologies and so on. But I’ll concentrate on the core models related to care delivery for the moment – terminology subsets and archetypes. These two form the basis for abstract formal modelling of content in a way that can be used to generate concrete expressions within implementation technologies like FHIR.
To make e-health a sustainable endeavour, we ideally need to have one set of domain content models and terminology subsets, and drive everything from that. This is the idea of the Clinical Information Modelling Initiative (CIMI), which aims to create a universal set of archetypes and terminology subsets primarily via importing definitions from Intermountain, openEHR, CDA templates, and any other model repositories. Within a CIMI model base there are essentially 2 levels:
- reference model (RM) archetypes – archetypes of generic information model concepts like ‘Observation’, ‘Assessment’ etc
- content archetypes – archetypes of real clinical content like ‘Adverse Reaction’.
There is also an underlying ‘CIMI Reference Model’, but this is trivial, and has no import on the semantics.
These two levels also exist in openEHR: the openEHR Reference Model and the openEHR archetypes; and in CDA as the CDA specification and CDA templates. CDISC has also been looking at moving to a ‘templated’ modelling approach, creating the same two levels.
There are also undoubtedly many health computing solutions built by vendors that a) are based on defined models of content and b) which do not treat their models of content as strategically proprietary, including some of the GP solutions found in the UK.
Realistically it will be some years before all of this gets funnelled through CIMI, not for lack of interest, but simply because in their current formats, these technologies already provide high business value, while converting everything to CIMI still requires a lot of work.
So now we have two challenges with respect to FHIR:
- how do we connect the world(s) of clinical content modelling to FHIR, making it more semantically sustainable?
- how do we connect FHIR to systems that are already model-based, as opposed to the non-model-based ones mentioned above?
An Extensibility Approach to FHIR
My proposition for solving these challenges is not the one currently assumed by most people. The usual assumption is that every implementer of a FHIR interface has to build a mapping from their underlying data to the FHIR published resources, primarily by ‘profiling’ and ‘extensions’. This is fine, if you have no explicit model of system content (or none that can be commercially exposed), but if you do, it’s a problem, because you are in the same position as trying to map your content to HL7v3 or any other message standard that wants to impose its own model of content – it’s costly and error-prone to do so.
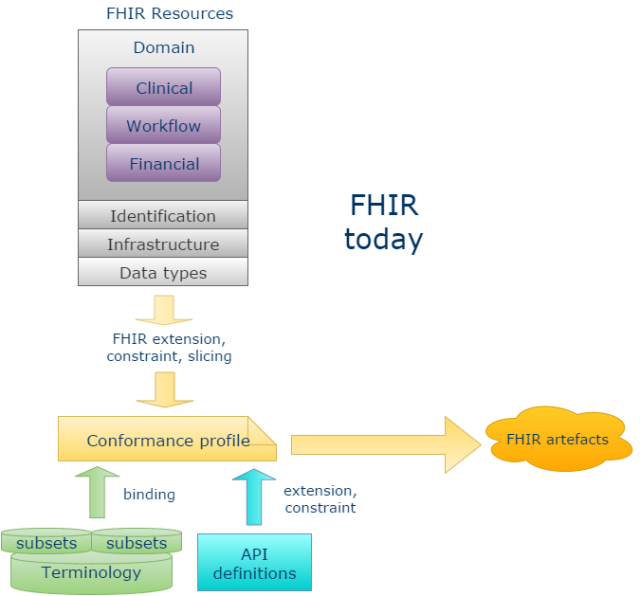
Doing this is the wrong approach for the model-based systems and architectures, in my view. There are actually two things that should be done differently. To understand them we need to consider FHIR in a different way than is currently done. The main change in thinking is to see FHIR as an API-building technology space rather than as a fixed API + content standard. This means that instead of thinking of a fixed set of resources, we should think of a ‘partitioned resource model space’, to which profiling and URI generation can be applied to create usable artefacts.
Concretely, we would stop thinking of the current FHIR clinical resources as some kind of definitive set, and instead think of them as a default set of fixed content models, for use with systems for which no published models are available. We should then assume that it is possible to add other resource ‘partitions’ to the resource space, enabling the plugging in of other model-based technologies to FHIR.
Secondly, clinical resource definitions in FHIR in the default set should be created based on models of content from sustainable and clinically-based sources like CIMI rather than built independently. This would ensure that users of those resources are connected to the mainstream clinical modelling activities in the world, and well-placed to use more modern model-based components based on those resources.
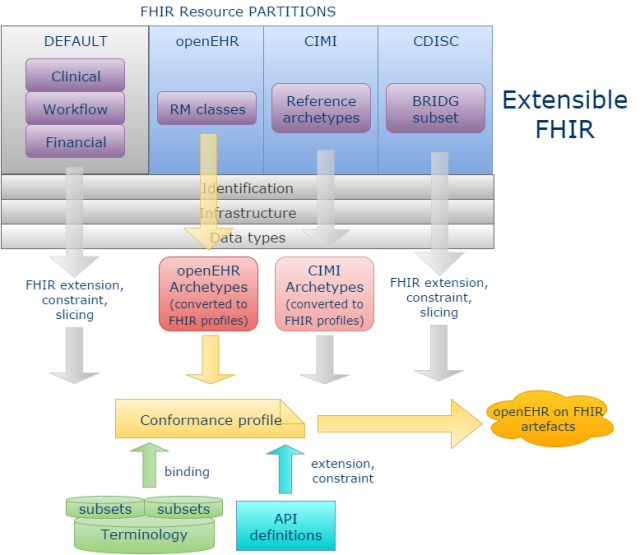
Below I have shown how this might work (with the development pathway for openEHR highlighted), by assuming that new FHIR resource partitions can be added for openEHR, CIMI and, to take an example from analytics, CDISC BRIDG. A new partition could be added for any architecture with published Reference Model and content models – including proprietary systems. I could have added a partition for CDA or CCDA as well, but have not done so, since my current assumption is that CDA users prefer to map to the FHIR default resources.
The partition in each case would contain FHIR resources that mimic the original ‘reference model entities’, but use the FHIR Data types, Identification and Infrastructure resources. The ‘Default’ partition contains the current FHIR Clinical, Workflow and Financial resources
For openEHR, these are just (some) Reference Model classes like Composition, Observation, Action etc; for CIMI, it will be mimics of the CIMI Reference Archetypes. For CDISC, it will be a subset of the BRIDG UML reference model.
We can now see in principle that openEHR clinical archetypes can be more easily mapped into FHIR, by converting them to FHIR profiles based on the openEHR partition resources, rather than the Default resources. Doing this also replaces the need to perform FHIR ‘extension, constraining and slicing’ activities, since the Archetype models have all this built-in. The same approach would be true of CIMI archetypes.
For BRIDG we would probably assume using the FHIR built-in extension, constraint, and slicing, but based on BRIDG resources rather than the default ones; an archetype approach could be used as well.
FHIR terminology and vanilla API definitions would be used across all model partitions.
The result of this way of doing things is that high-quality FHIR artefacts can be generated for model-based architectures, that are known to be faithful to their underlying semantic models, but also to the FHIR infrastructure and identifier resources, data types, and terminology. This could greatly extend the utility of FHIR from being limited to profiles on a fixed set of resources – not that different to a fixed set of messages, if we think about it – to working for any architecture.
Is there a downside? Well, clearly it’s a more ambitious project, but on the other hand, a huge amount of mapping work evaporates, and quality of output artefacts should be better than if the usual manual mapping approach had been attempted.
The other downside is that a CIMI or openEHR FHIR blood pressure won’t look exactly like a BP based on the default FHIR Observation resource. However, if a set of vital signs resources were required that worked the same way across all FHIR resources, it could be addressed with some mapping within the FHIR model space where all of the data types, infrastructure and terminology problems have already been dealt with, rather than trying to perform complex mappings across completely different model spaces.
Based on 15 years experience with trying to map various EHR and message architectures to each other, there is no realistic hope anyway of achieving a single FHIR resource for every clinical content entity that works for everyone, if it is mandatory to use the default set of FHIR resources, because this is just the same complex mapping problem we already know can’t be solved in a general way. So we need a different approach.
The above proposal won’t change anything for many people already working on FHIR profiles based on the default Clinical and other resources, but would potentially provide a good solution for communities like CIMI, openEHR, 13606, CDISC and many others to use FHIR without having to fight with difficult mappings.
Naturally in the above I have glossed over many details which would need to be sorted out to make the proposal a going concern. But first I think we need to see if the vision excites anyone’s interest.
[See this post at HealthIntersections.com for more discussion.]


It could just be that there is not an appetite for internationally agreed clinical models. Most of the projects I work on the clinicians are focussing on local needs or at best national needs, they seldom care about the new international view. It is the informaticians who try to sell this view not the clinicians or indeed most of the systems vendors.
From what I have seen of FHIR it is maturing really quite quickly. The models seems to be more intuitive and understandable to clinicians as they more closely represent real world objects than other approaches which are more abstract.
My thoughts are that if we can get a collaborative review tool in place for FHIR along the same lines of CKM for openEHR then that will make a tremendous difference.
The proliferation of the ecosystem around FHIR with various tools and code libraries is impressive and growing both in breadth and maturity. This is proportional to the rate of implementation and adoption by the multinational vendors and national programs alike. FHIR is still young, I suspect that in another year of two then we will see large scale implementations around the world.
Fundamentally there has to be a reason why other methodologies have not achieved this and FHIR IS doing so….
Just on the topic of ‘international’ versus ‘national’ etc.. here in Europe, ‘international’ just means an activity that may have had input from England, Scotland; it’s pretty common for some level of cooperation to exist across EU member countries. ‘National’ is the main focal level for most content modellers – each country has its own needs and priorities. If clinical people only model locally it will be a problem for obvious reasons. If FHIR is implemented on a local basis, it won’t matter much, because FHIR is about querying, not about data capture.
I don’t think it’s as clear cut as that. Colleagues inform me that the English pathology work is looking to align closely with the US work in this area for example. FHIR is also far more than query, though admittedly this is the low hanging fruit that early implementations are focussing on currently. The FHIR initiatives around workflow and questionnaire are very much more than query.
This looks like using FHIR as little more than a syntax/transport layer, and pushes the mapping activity into the CIMI / openEHR layer (because you still have to bridge from a native system). Furthermore, claiming that mapping “within the FHIR model space” is somehow easier seems like the same fallacy perpetrated by some people about XML/RDF/… that once everything is in the same form, then the mapping is easier. It’s not – the (hard) mismatches are at the conceptual modelling level and remain.
It also looks somewhat like an us vs them approach in that it turns its back on the existing clinical Resources rather than looking to embrace and improve them.
What I’d be interested in understanding is why, for example, the openEHR / CIMI blood pressure model can’t be mapped on top of (ie as an extension to) the BP in a FHIR Observation, given that FHIR should be capturing “the common 80%”
Finally, what I’m also missing here is the motivation for wanting to do this in the first place (not saying there aren’t good reasons) — what is the problem that this solves and the value that it provides?
The real difficulties in mapping are indeed conceptual (I have commented many times myself to the same effect). However, I am assuming that FHIR can be treated not just as a literal syntax like RDF or XML etc, but that it has some semantics of its own that everyone can agree on – data types, infrastructure resources, use of terminology, approach to generating URIs for an API and so on. If an architecture with its own model space can map its main semantics into FHIR while converting the above-mentioned things to FHIR-form, part of the problem is solved.
But in the end I’m not that worried by trying to make complete mappings across major architectures – it doesn’t happen today, and it’s only needed in a few cases anyway – vital signs, the main managed lists etc.
Take the BP example – it’s easy enough to map a single ‘systolic BP’ from openEHR to FHIR. But if you want to map a 1000 samples in time, each containing a particular structure of systolic, diastolic, cuff size, exertion, instrument, and a particular arrangement of context attributes, it’s not easy at all. And yet this is absolutely routine data. Now, natively in openEHR, FHIR isn’t needed to expose this kind of data – the FHIR equivalent layer is mostly generated from the archetypes – see https://www.ehrscape.com/api-explorer.html
But if the openEHR RM classes existed as FHIR resources, things change completely. Then anyone using the FHIR infrastructure can easily get to even the most complicated openEHR (or CIMI, CDISC, proprietary) data in a natural way.
Re: us v them, well that argument can apply in either direction. In openEHR there are 50 times the number of clinical resources of FHIR; if we include CIMI/Intermountain, it becomes 100. Plus the rate of creation of more archetypes is a lot higher. There’s no realistic option for openEHR to rely on FHIR’s clinical resources. That’s not a criticism of the latter – it’s just the way things actually are.
I would have thought that some people working in the FHIR space might be interested in ways to bringing that much larger catalogue of clinical models into FHIR, other than by manual re-invention. At least that’s my hope.
What stays is that FHIR is a format/method to exchange data between EXISTING databases in an defined Interface.
It does not extend to full semantic interpretability in a sense that future systems, clinical decision support systems, future researchers can interpret the data safely without an implementation guide and human intervention.
The FHIR defined data is NOT safely semantically interpretable by a computer taking the complete context in consideration.
FHIR does define computable information models and their terminology bindings for a limited number of administrative and clinical concept. Thus it is as good in terms of semantics as any other content standard. Plus it provides the means to add more stuff by extensions and slicing/profiling mechanisms – which are all computable and referencable. Therefore it is as computable and semantically ‘interpretable’ as openEHR or 13606. The big difference is it lacks the dual-level or multi-level modelling method – that is it is ‘single-level’ modelling and hence lacks the benefits of true model-driven development approach. But one should keep in mind that it is not engineered for system development; e.g. persistence and UI etc. As an HIE formalism I think there has been a deliberate and inevitable trade-off between being ‘implementer-friendly’ and ‘clinician/business user-friendly’. The real problem is FHIR has some 50 resources now and it took X amount of time and effort, the next 50 resources will take >2X – probably will grow rather exponentially due to the need to keep resources (plus downstream extensions+profiles) consistent and clinically valid/safe. I strongly think openEHR can complement FHIR by providing much needed clinical content so they can continue to focus on ‘implementer-friendly’. I’m very hopeful that it won’t be US vs. rest of the world this time as most people I know are quite sensible and intelligent plus vendors & governments are putting a lot of pressure to get some real benefits. So the time is ripe for win-all.
My 2 cents
Great post – thanks Thomas.
Regarding your comment on CDA, “I could have added a partition for CDA or CCDA as well, but have not done so, since my current assumption is that CDA users prefer to map to the FHIR default resources.”, please note that there are two efforts: (1) CDA on FHIR and (2) CCDA FHIR profiles (direct) development; the former is more similar I believe to the approach you suggest in your post for openEHR, CIMI and BRIDGE.
Also, you wrote “These two levels also exist in openEHR: the openEHR Reference Model and the openEHR archetypes; and in CDA as the CDA specification and CDA templates. CDISC has also been looking at moving to a ‘templated’ modelling approach, creating the same two levels.” – we have to keep in mind that CDA is built on the HL7 RIM, so there is another layer in this case. However, what CDA adds on top of the RIM is its clinical statement model, which is based on the allowed associations between the HL7 RIM classes. While core classes of most reference models are quite similar (observation, procedure, medication, etc.), the model of how they can be associated with each other is still a challenge.